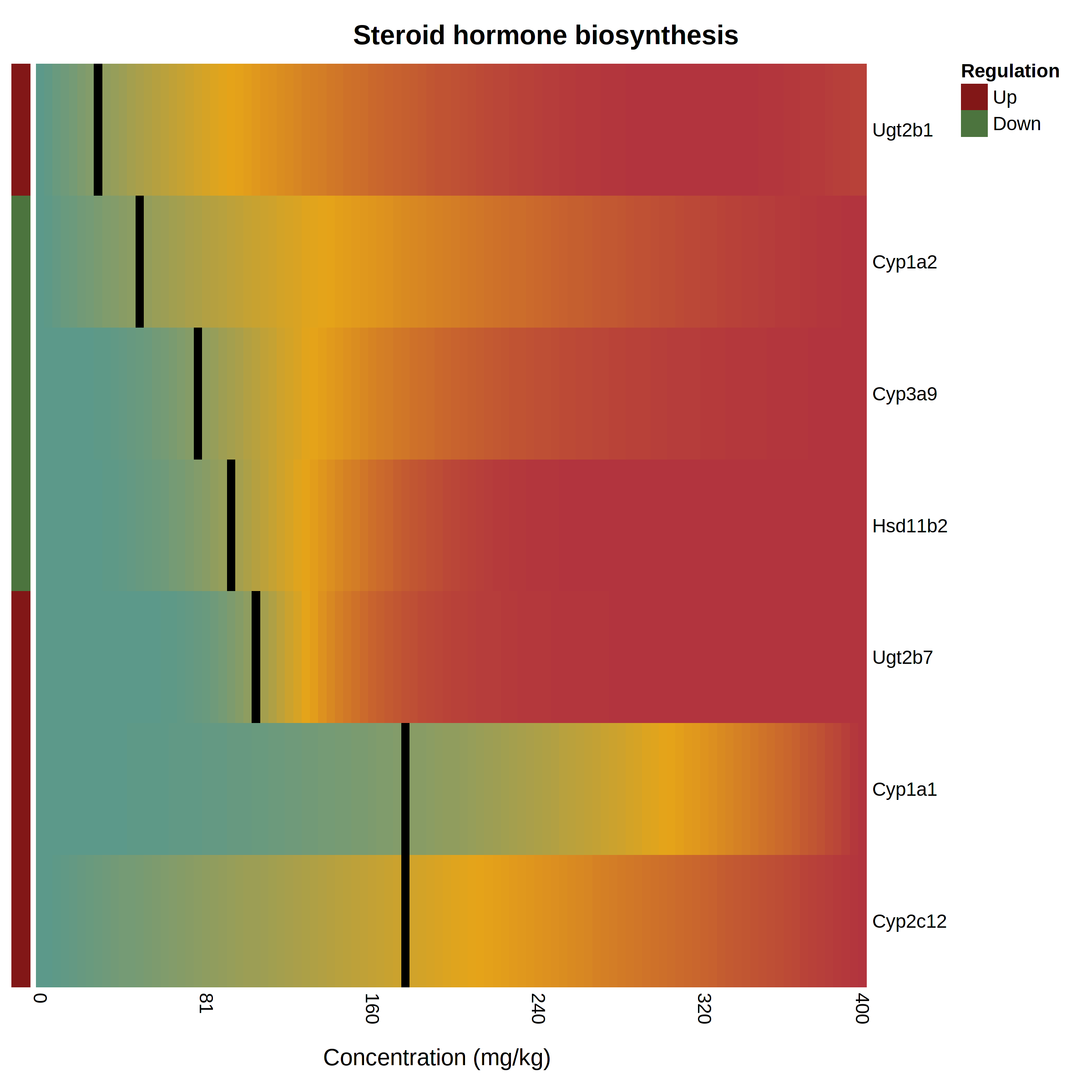
The pathway heatmap is an appealing visualization to clearly shows how the expression of each pathway gene compares to the others. It is generated when you click a pathway or gene set name in the “Gene Set Enrichment” panel at the result page. An example output is shown below The pathway heatmap values are calculated through a series of steps: The fitted model for each gene is evaluated across the range of doses in the uploaded data. The resulting modeled expression values are normalized
Heatmap - an overview – Flourish
Heatmap - an overview – Flourish

The heatmap and volcano plot of DEGs. (a) In the heatmap, red color

Transcriptome and metabolome profiling provides insight into the regulatory network of fruit coloration in Coffea arabica L - ScienceDirect
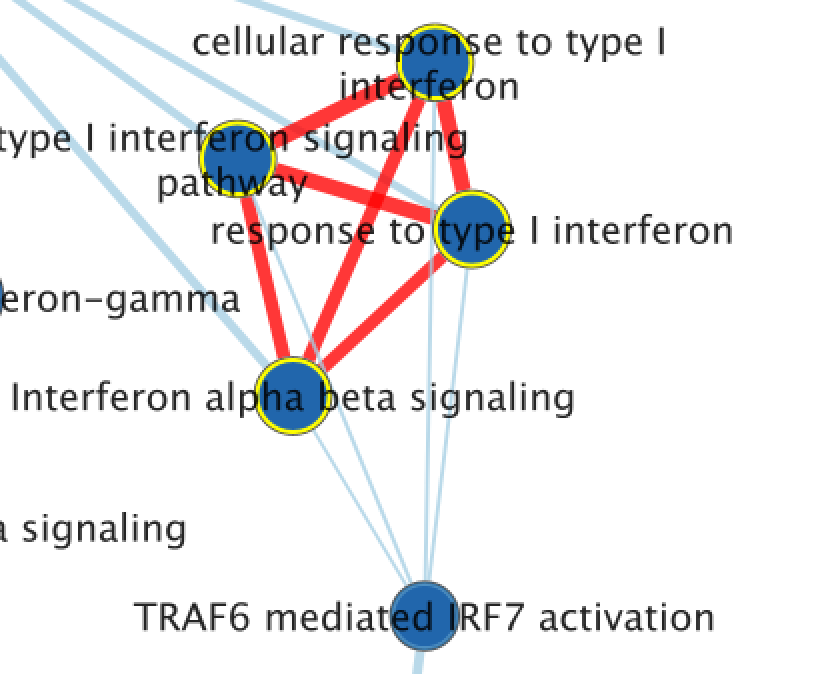
13A) Exploring the Table Panel heat map Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap
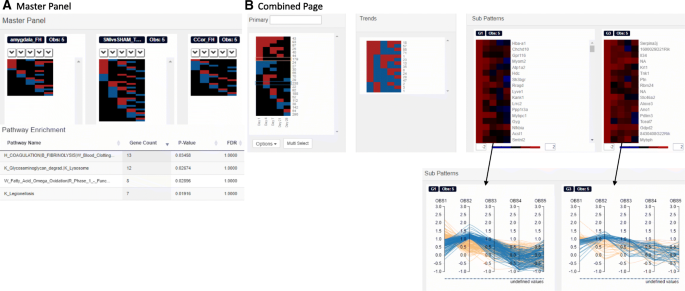
Functional Heatmap: an automated and interactive pattern recognition tool to integrate time with multi-omics assays, BMC Bioinformatics
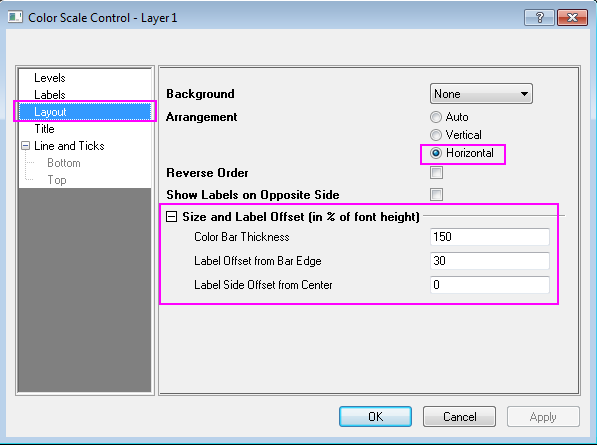
d2mvzyuse3lwjc.cloudfront.net/doc/en/Tutorial/imag

The heatmap and clustering analysis for metabolic pathways. a Heatmap

Bitesize Bioinformatics: Drawing Heatmaps in R
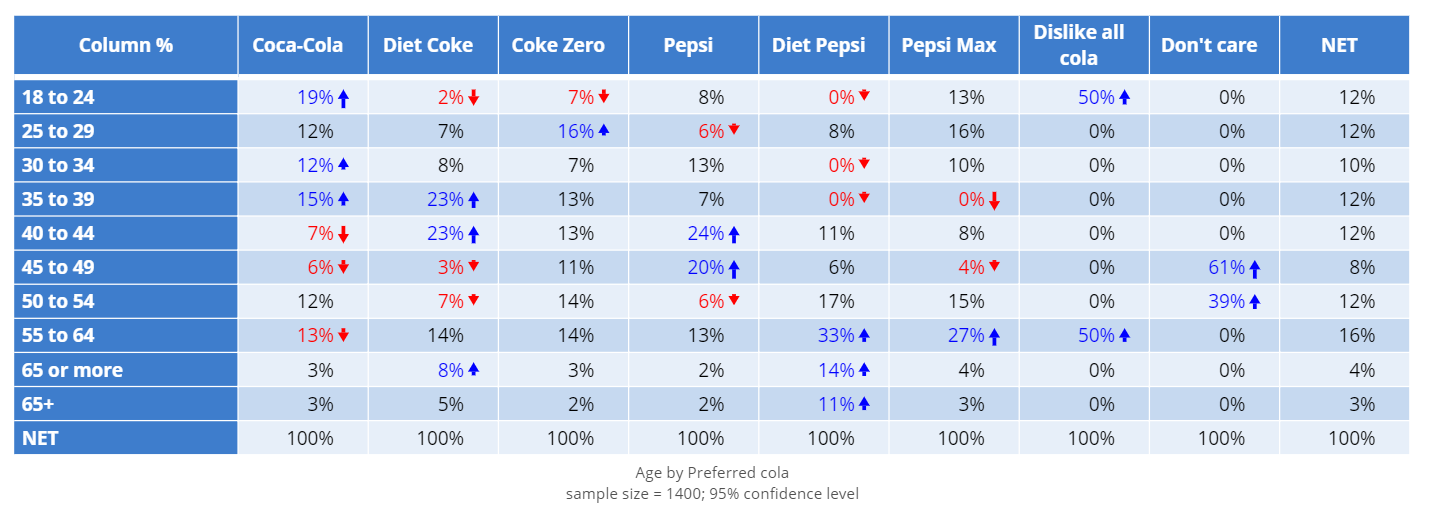
Visualization - Heatmap - Heatmap - Q